Summary
General-purpose AI models like Segment Anything (SAM) and CellPose have transformed computer vision and brought powerful new capabilities into the life sciences. But real microscopy data is messy: imaging conditions and quality vary, staining intensity differs, debris and dust appear, and various assays demand very distinct readouts. This article reviews why AI cell segmentation with universal models often fails in practice, how these limitations affect reproducibility, and why the field is moving toward custom, assay-specific models. We highlight examples from academia, note “train-your-own” solutions, and discuss how AI-as-a-service platforms, such as SnapCyte™, lower barriers for adoption.
The Promise of Universal Segmentation Models
Over the past few years, general-purpose AI cell segmentation tools have generated enormous excitement. Meta’s SAM, trained on over a billion masks, demonstrated the feasibility of “zero-shot” segmentation across diverse domains, including microscopy. Similarly, CellPose, trained on a large and varied set of biological images, became one of the first practical community tools for cell segmentation (Stringer et al., 2021). Omnipose further improved performance for elongated and irregularly shaped cells (Cutler et al., 2022).
These tools reduce reliance on hand-tuned thresholds, macros, and subjective manual annotation. On clean, high-quality images at standard magnifications, they can exceed human performance and deliver reproducible results—an important step forward compared to traditional threshold-based image processing.
Why Real Microscopy Data Doesn’t Fit Generalization
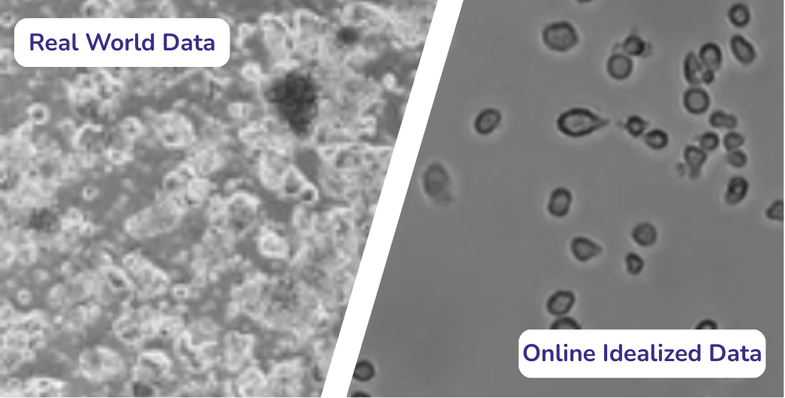
In practice, however, real microscopy data rarely looks like training data. Images often contain debris, contamination, autofluorescence, scratches or uneven illumination. Even advanced AI cell segmentation models trained on curated datasets frequently struggle in these everyday contexts.
Instrument variability: Imaging parameters (objective, camera, illumination, magnification) strongly affect appearance, and cross-site domain shifts are a known source of error (Stacke et al., 2020).
Assay heterogeneity: Different assays require different outputs. Various cells add further complexity, with mixed morphologies and overlapping structures.
Sample diversity: 2D cell culture, 3D structures, tissues, spheroids, and tissue slides exhibit 3D organization and irregular boundaries. Models trained on monolayers and uniform sample types often over- or under-segment these systems (Boutin et al., 2018).
Staining and contrast: Bright field images have low contrast and strong background variation, while fluorescence markers differ by channel, background signal and noise. The LIVECell dataset highlighted how generalist models lose accuracy on low-contrast, high-density brightfield images (Edlund et al., 2021).

The Reproducibility Gap in Microscopy
Scientists increasingly demand reproducibility across experiments, labs, and operators. Manual workflows, even when supported by “universal” AI cell segmentation models, often leave room for bias:
Thresholds and parameters are adjusted by individual researchers.
Borderline cases (e.g., overlapping cells, low-contrast images) generate inconsistent results.
Cross-lab comparisons rarely match because imaging conditions differ.
This reproducibility gap explains why many labs still default to manual data analysis, despite the availability of large pretrained tools.
Custom AI Models as the Practical Path Forward
Custom models address these limitations by adapting strong backbones (SAM, CellPose, Omnipose, transformers) to assay-specific datasets. Instead of forcing a general model to fit every case, researchers fine-tune it for their exact experimental setup and imaging conditions.
Benefits include:
Consistency: Every image is analyzed against the same criteria, reducing operator-to-operator variation.
Assay-specific outputs: Models can be tailored for quantitative readouts beyond segmentation.
Scalability: Once validated, the same model can process hundreds or thousands of images without further tweaking.
Microscope vendors such as Zeiss (via arivis AI) and Olympus/Evident (TruAI) already recognize the limits of universal segmentation and now offer “train-your-own-model” options. These mark an important shift: instead of forcing one universal algorithm to work everywhere, they allow labs to create smaller, assay-specific AI cell segmentation models.
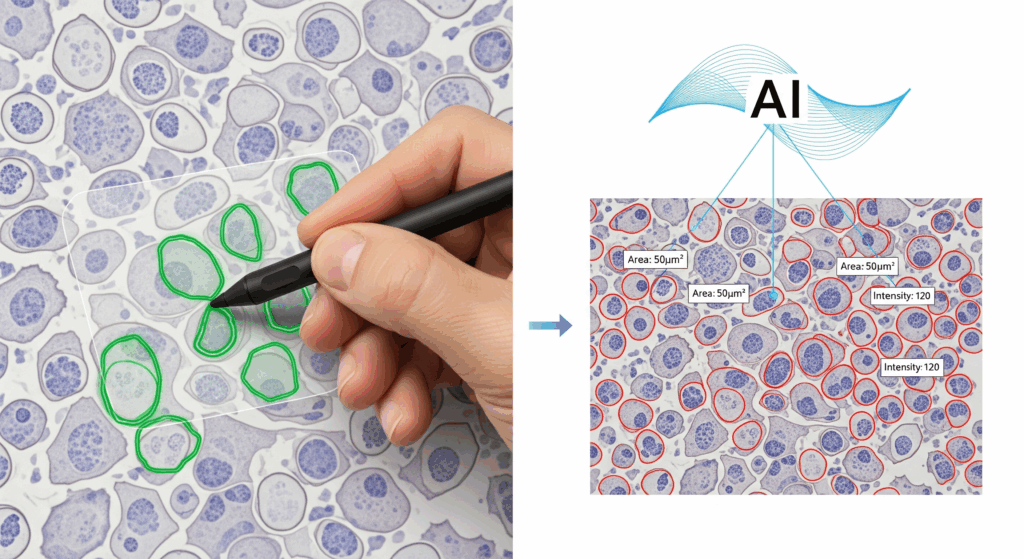
In practice, though, the biggest bottleneck is annotation. Creating ground-truth masks requires meticulous work: drawing precise boundaries, separating overlapping cells, filtering debris, and making judgment calls in ambiguous cases. Faintly stained cells, partially out-of-focus objects, or overlapping morphologies often force researchers into subjective decisions. Without clear guidelines, annotations differ between users, leading to biased or inconsistent training sets.
Even trained experts can struggle. A study in computational pathology showed that noisy labels and inconsistent annotations directly undermine model reliability, producing classifiers that replicate biases instead of resolving them [Marzahl et al., 2020]. Microscopy poses the same risk: scientists are rarely trained in annotation best practices, and quality-assurance steps are often skipped. In edge cases, such as deciding whether a dim cell is on the same plane or not, scientists may not agree on the “correct” answer.
Vendor platforms provide the interface, but they leave annotation, quality control, and scientific interpretation entirely to the lab. This is precisely where adoption breaks down: the tools are powerful, but they demand time, expertise, and infrastructure most labs don’t have.
AI-as-a-Service: Lowering the Barrier
AI-as-a-service offers a different approach. Instead of asking every lab to annotate and train, service providers take over the entire pipeline.
Labs can start with prebuilt models for common assays (e.g., cell counting, confluency) to get results immediately.
For custom workflows, providers like SnapCyte™ work directly with researchers to define the biological readouts that matter.
Annotation and quality assurance are handled centrally, using consistent guidelines and validated pipelines, so the resulting models aren’t biased by inconsistent local labeling.
Once trained, the model is deployed back into a ready-to-use platform that works across imaging modalities.
This means scientists get outputs aligned with their experiments, without needing to code, manage plug-ins, or retrain models every time conditions shift.
Final Thoughts
Generalist models are remarkable achievements, but they cannot fully bridge the gap between curated benchmarks and the messy, variable reality of microscopy. Annotation bottlenecks, reproducibility issues, and assay heterogeneity make it clear that custom, assay-specific AI cell segmentation approaches are the practical path forward. Vendor solutions have started to move in this direction, but adoption remains limited because they leave annotation and quality control to end users.
AI-as-a-service represents the next wave: reproducible models trained on real lab data, deployed in platforms that scientists can use immediately, without technical overhead.
👉 If you’d like to explore how SnapCyte can support your assays with prebuilt or custom models, get in touch with us.
References
- Segment Anything Model (SAM) – Meta AI, 2023
- Stringer, C., Wang, T., Michaelos, M. et al. Cellpose: a generalist algorithm for cellular segmentation. Nat Methods 18, 100–106 (2021). https://doi.org/10.1038/s41592-020-01018-x
- Cutler, K.J., Stringer, C., Lo, T.W. et al. Omnipose: a high-precision morphology-independent solution for bacterial cell segmentation. Nat Methods 19, 1438–1448 (2022). https://doi.org/10.1038/s41592-022-01639-4
- Stacke K, Eilertsen G, Unger J, Lundstrom C. Measuring Domain Shift for Deep Learning in Histopathology. IEEE J Biomed Health Inform. 2021 Feb;25(2):325-336. doi: 10.1109/JBHI.2020.3032060. Epub 2021 Feb 5. PMID: 33085623.
- Boutin, M.E., Voss, T.C., Titus, S.A. et al. A high-throughput imaging and nuclear segmentation analysis protocol for cleared 3D culture models. Sci Rep 8, 11135 (2018). https://doi.org/10.1038/s41598-018-29169-0